Add 500 μl wash buffer and then wash 2x with 1 ml high-salt buffer.Carefully remove the supernatant and resuspend the beads in 20 μl ligation mix (9 μl water 4 μl 4x ligation buffer 1 μl RNA ligase 0.5 μl RNasin 1.5 μl pre-adenylated linker 元 4 μl PEG400 ).Add 500 μl wash buffer and wash 1x with high-salt buffer.Discard the supernatant and resuspend the beads in 20 μl PNK mix (15 μl water 4 μl 5x PNK pH 6.5 buffer 0.5 μl PNK enzyme 0.5 μl RNasin ).Wash 2x with 900 μl wash buffer (20 mM Tris-HCl, pH 7.4 10 mM MgCl 2 0.2% Tween-20).Discard the supernatant and wash the beads 2x with 900 μl high-salt buffer (50 mM Tris-HCl, pH 7.4 1 M NaCl 1 mM EDTA 1% NP-40 0.1% SDS 0.5% sodium deoxycholate). 
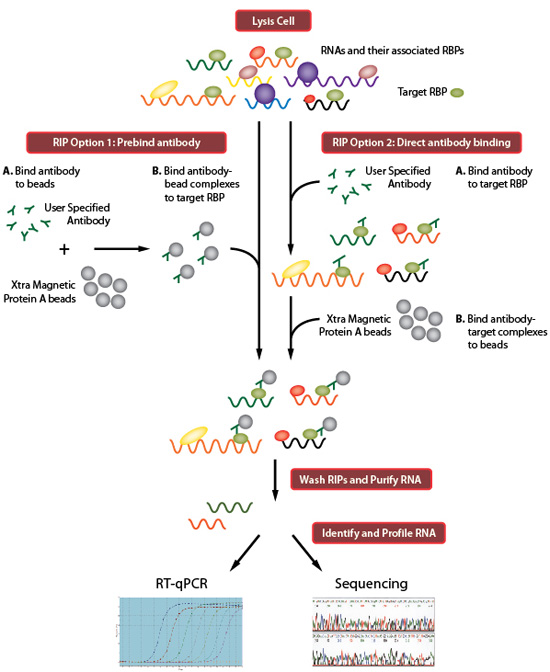
Remove the wash buffer from the beads (from step 2.5), then add the cell lysate (from step 3.4).Carefully collect the supernatant (leave about 50 μl lysate with the pellet). Spin at 4☌ and 22,000 g for 20 min to clear the lysate.Incubate the samples for exactly 3 min at 37☌, shaking at 1,100 rpm.Add 10 μl RNase I dilution as well as 2 μl Turbo DNase to the cell lysate (1/500 RNase I dilutions are used for library preparation 1/50 dilutions are necessary to control for antibody specificity). Prepare a 1/500 dilution of RNase I (Ambion, AM2295).Resuspend the cell pellet in 1 ml lysis buffer and transfer to 1.5 ml microtubes.Wash 3x with 900 μl lysis buffer and leave in the last wash until ready to proceed to step 4.1.Rotate tubes at room temperature for 30-60 min.Resuspend beads in 100 μl lysis buffer with 2-10 μg antibody.Wash beads 2x with lysis buffer (50 mM Tris-HCl, pH 7.4 100 mM NaCl 1% NP-40 0.1% SDS 0.5% sodium deoxycholate 1/100 protease inhibitor cocktail III, Calbiochem).Add 100 μl of protein A Dynabeads (Dynal, 100.02) per experiment to a fresh microtube (Use protein G Dynabeads for a mouse or goat antibodies).Snap-freeze the cell pellets on dry ice and store at -80☌ until use.Spin at top speed for 10 sec at 4☌ to pellet cells, then remove supernatant. Transfer 2 ml cell suspension to each of three microtubes.Harvest the cells by scraping with a cell lifter.Irradiate once with 150 mJ/cm 2 at 254 nm. Remove the media and add 6 ml ice-cold PBS to cells grown in a 10 cm plate (enough for three experiments).We successfully applied iCLIP to study hnRNP C particle organization on a genome-wide scale and assess its role in splicing regulation 14.ġ. Importantly, sequencing the truncated cDNAs provides insights into the position of the cross-link site at nucleotide resolution. We recently developed iCLIP (individual-nucleotide resolution CLIP), which captures the truncated cDNAs by replacing one of the inefficient intermolecular RNA ligation steps with a more efficient intramolecular cDNA circularization (Figure 1) 14. Such truncated cDNAs are lost during the standard CLIP library preparation protocol. In addition, primer extension assays indicated that many cDNAs truncate prematurely at the crosslinked nucleotide 13. This is partly due to the restricted amount of co-purified RNA and the two inefficient RNA ligation reactions required for library preparation. Recently, PAR-CLIP was introduced that uses photoreactive ribonucleoside analogs for cross-linking 11,12.ĭespite the high specificity of the obtained data, CLIP experiments often generate cDNA libraries of limited sequence complexity. In combination with high-throughput sequencing technologies, CLIP has proven as a powerful tool to study protein-RNA interactions on a genome-wide scale (referred to as HITS-CLIP or CLIP-seq) 9,10. CLIP combines UV cross-linking of proteins and RNA molecules with rigorous purification schemes including denaturing polyacrylamide gel electrophoresis. In order to increase the specificity and positional resolution, a strategy referred to as CLIP (UV cross-linking and immunoprecipitation) was introduced 7,8. These approaches were prone to identifying indirect or non-physiological interactions 6. Initial attempts to study protein-RNA complexes in their cellular environment employed affinity purification or immunoprecipitation combined with differential display or microarray analysis (RIP-CHIP) 3-5. Protein-RNA interactions can be studied using biochemical methods, but these approaches do not address RNA binding in its native cellular context. Therefore, an essential step towards understanding transcript regulation at the molecular level is to gain positional information on the binding sites of RBPs 2. The unique composition and spatial arrangement of RNA-binding proteins (RBPs) on a transcript guide the diverse aspects of post-transcriptional regulation 1.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
